Quickstart
A Bayesian Belief Network (BBN) is defined as a pair \((D, P)\) where
\(D\) is a directed acyclic graph (DAG), and
\(P\) is a joint distribution over a set of variables corresponding to the nodes in the DAG.
Creating a reasoning model involves defining \(D\) and \(P\). In py-scm, the current reasoning surface is exact for linear-Gaussian causal models defined by a DAG, a mean vector, and a covariance matrix.
Associational: conditional Gaussian queries.
Interventional: exact post-intervention queries under the \(\mathrm{do}\) operator.
Counterfactual: exact abduction-action-prediction queries when the factual world is fully observed.
In this notebook, we show how to create a Gaussian BBN and conduct the different types of causal inference supported by py-scm.
Creating a model
Create the structure, DAG
Creating the DAG means to define the nodes and directed edges.
[1]:
d = {"nodes": ["C", "X", "Y"], "edges": [("C", "X"), ("C", "Y"), ("X", "Y")]}
[2]:
from pyscm.serde import dict_to_graph
import networkx as nx
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(5, 5))
g = dict_to_graph(d)
pos = {"C": (0, 1), "X": (1, 0), "Y": (2, 1)}
nx.draw(g, pos=pos, with_labels=True, node_color="#e0e0e0", ax=ax)
fig.tight_layout()
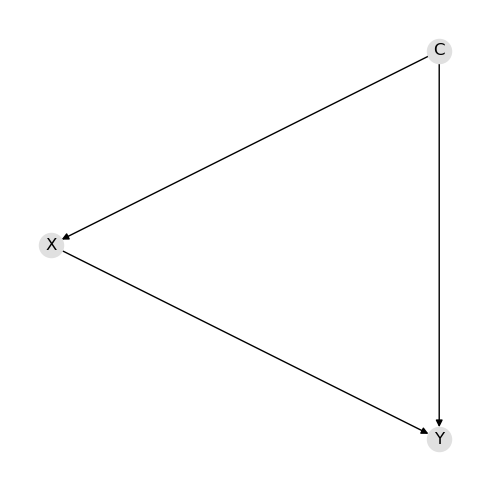
Create the parameters
Creating the parameters means to define the means and covariance matrix. The means and covariance below were estimated from the following normal distributions.
\(C \sim \mathcal{N}(1, 1)\)
\(X \sim \mathcal{N}(2 + 3 C, 1)\)
\(Y \sim \mathcal{N}(0.5 + 2.5 C + 1.5 X, 1)\)
[3]:
p = {
"v": ["C", "X", "Y"],
"m": [1.00172341, 4.99599921, 10.5032959],
"S": [
[0.99070024, 2.97994442, 6.95690224],
[2.97994442, 9.97338239, 22.44685389],
[6.95690224, 22.44685389, 52.12803651],
],
}
Create the model
Finally, we can create the reasoning model once we define the DAG and parameters.
[4]:
from pyscm.reasoning import create_reasoning_model
model = create_reasoning_model(d, p)
Associational query
You are able to conduct associational query with and without evidence.
Query without evidence
Associational query involves invoking the pquery() method. A tuple is returned where the first element is the means and the second element is the covariance matrix. The means and covariance matrix are the parameters of the multivariate normal distribution.
[5]:
q = model.pquery(pandas=True)
[6]:
q[0]
[6]:
C 1.001723
X 4.995999
Y 10.503296
dtype: float64
[7]:
q[1]
[7]:
| C | X | Y | |
|---|---|---|---|
| C | 0.990700 | 2.979944 | 6.956902 |
| X | 2.979944 | 9.973382 | 22.446854 |
| Y | 6.956902 | 22.446854 | 52.128037 |
Query with evidence
If you have evidence, pass in a dictionary of the observed evidence to pquery().
[8]:
q = model.pquery({"X": 2.0}, pandas=True)
[9]:
q[0]
[9]:
C 0.106550
X 2.000000
Y 3.760272
dtype: float64
[10]:
q[1]
[10]:
| C | X | Y | |
|---|---|---|---|
| C | 0.100323 | 2.979944 | 0.250012 |
| X | 2.979944 | 9.973382 | 22.446854 |
| Y | 0.250012 | 22.446854 | 1.607438 |
Interventional query
Interventional query involves graph surgery where the incoming edges to the manipulated variable are removed. Interventional query is conducted by invoking the iquery() method. Below, the \(\mathrm{do}\) operation is applied to \(X\). The method returns a series with the post-intervention mean and std for the target variable.
[11]:
model.iquery("Y", {"X": 2.0}, pandas=True)
[11]:
mean 5.991103
std 2.671521
dtype: float64
Compare queries
Let’s compare the results of the associational and interventional queries by using the resulting parameters to sample data.
[12]:
import pandas as pd
pd.Series(
{
"marginal": model.pquery(pandas=True)[0].loc["Y"],
"conditional": model.pquery({"X": 2.0}, pandas=True)[0].loc["Y"],
"causal": model.iquery("Y", {"X": 2.0}, pandas=True).loc["mean"],
}
)
[12]:
marginal 10.503296
conditional 3.760272
causal 5.991103
dtype: float64
Counterfactual
Counterfactual queries are conducted using cquery(). You must pass the factual evidence and one or more counterfactual interventions. The factual assignment must include every variable in the model so that the abduction step is fully determined. The result is a dataframe containing the target parents used in each hypothetical world together with the factual and counterfactual values for the target.
Below, the factual evidence, f, is what has already happened. The counterfactual interventions, cf, are the hypothetical actions we want to evaluate. In plain language, we are asking the following.
Given we have observed C=0.945536, X=4.970491, Y=10.542022, what would have happened to Y if
X=1?
X=2?
X=3?
C=2 and X=3?
[13]:
f = {"C": 0.945536, "X": 4.970491, "Y": 10.542022}
cf = [{"X": 1}, {"X": 2}, {"X": 3}, {"C": 2, "X": 3}]
q = model.cquery("Y", f, cf, pandas=True)
[14]:
q
[14]:
| C | X | factual | counterfactual | |
|---|---|---|---|---|
| 0 | 0.945536 | 1.0 | 10.542022 | 4.562174 |
| 1 | 0.945536 | 2.0 | 10.542022 | 6.068247 |
| 2 | 0.945536 | 3.0 | 10.542022 | 7.574319 |
| 3 | 2.000000 | 3.0 | 10.542022 | 10.202112 |
Data sampling
To sample data from the model, invoke the samples() method.
[15]:
sample_df = model.samples()
sample_df.shape
[15]:
(1000, 3)
[16]:
sample_df.head()
[16]:
| C | X | Y | |
|---|---|---|---|
| 0 | 1.899734 | 7.130419 | 16.506616 |
| 1 | 0.313851 | 3.478815 | 6.510815 |
| 2 | -0.100981 | 0.740561 | 2.555726 |
| 3 | 1.470743 | 7.623403 | 15.504343 |
| 4 | 0.999349 | 5.272057 | 10.237443 |
[17]:
sample_df.mean()
[17]:
C 0.965382
X 4.906904
Y 10.259403
dtype: float64
[18]:
sample_df.cov()
[18]:
| C | X | Y | |
|---|---|---|---|
| C | 1.020976 | 3.101065 | 7.238579 |
| X | 3.101065 | 10.416816 | 23.446800 |
| Y | 7.238579 | 23.446800 | 54.473860 |
Serde
Saving and loading the model is easy.
Serialization
To persist the model, use the model_to_dict() method.
[19]:
import json
import tempfile
from pyscm.serde import model_to_dict
data1 = model_to_dict(model)
with tempfile.NamedTemporaryFile(mode="w", delete=False) as fp:
json.dump(data1, fp)
file_path = fp.name
print(f"{file_path=}")
file_path='/tmp/tmpthmc6yt9'
Deserialization
To load the model, use the dict_to_model() method.
[20]:
from pyscm.serde import dict_to_model
with open(file_path, "r") as fp:
data2 = json.load(fp)
model2 = dict_to_model(data2)
[21]:
model2
[21]:
ReasoningModel[H=[C,X,Y], M=[1.002,4.996,10.503], C=[[0.991,2.980,6.957]|[2.980,9.973,22.447]|[6.957,22.447,52.128]]]